TECHNOLOGY
The lead that New Amsterdam Genomics has in the marketplace is due to our technology. We have made significant strides in the analysis of DNA data, enabling far more people to receive value from getting their DNA sequenced.
DNA Sequencing
We specialize in whole exome and whole genome sequencing, which provides much wider coverage than traditional genotyping or gene panel services. The exome is the entire protein-coding region of the human genome and includes over 22,000 genes. While comprising just 1% of our total DNA, the exome encompasses the vast majority of disease-causing mutations. The exome is generally the most effective and efficient genetic analysis for the clinical diagnosis, prognosis, and optimal treatment of disease.
Below is a diagram showing genotyping (in red), which looks at DNA at just a few places; gene panels (in green), which looks at just a few genes of DNA; and whole exome (in blue), which examines all 22,000 genes.
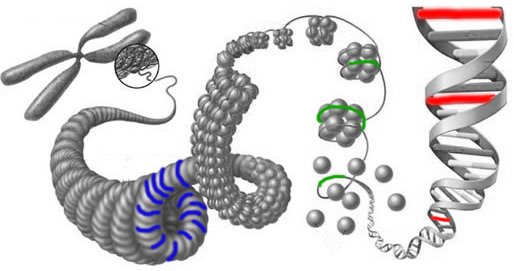
Next Generation Sequencing (NGS) is an enabling technology that has dramatically lowered the cost of DNA sequencing. As part of NGS, DNA is spliced into small fragments (usually around 100 nucleotides each, but it can be longer) and then sequenced in parallel using a fluorescence-based biochemical technique. The sequence output for each of these fragments is referred to as a read, and these raw reads go through a rigorous quality control process where adapters and sequences of low confidence are discarded.
DNA Analysis
We differentiate ourselves by our powerful computer algorithms to detect and interpret genomic variation.
First, we compare each ~100 base-pair processed read to what is called the reference genome. This process is analogous to a puzzle, where the reference genome is like the picture on the front of the box and the 100 base-pair sequences are the puzzle pieces. Each section of the genome must be covered by about 100 different reads to provide clinical-quality confidence. After the mapping algorithms have reconstructed the person's genome from the fragments, the next step is analysis.
By comparing the person's reads to the reference genome, we determine which variants the person has. There are many different types of variants: single point mutations, small insertions and deletions, and structural variants like copy number variations and chromosomal rearrangements. We focus on those variants that we can confirm with the highest accuracy.

Interpretation and Meaning
After deciding which mutations a person has, our algorithms then determine their impact. These interpretation algorithms are an essential reason N.A.G. is able to deliver much more value than other DNA tests. Some components of these algorithms include:
- constructing biological models to determine the impact of a mutation
- performing natural language processing of published papers to find mentions of a mutation to understand what these papers assert
- aggregating all the information about a single mutation and deriving conclusions
- aggregating all the conclusions about all the individual mutations related to a particular disease to determine an overall polygenic risk score for that disease
- determining how genetic risks interact with reported lifestyle factors and lab measurements to quantify a person's overall disease risks
Going Deep
In the case that a patient has mysterious symptoms or a family history of cancer, we go deep. No other organization is willing or even able to investigate a patient as deeply as us. We have developed novel computer algorithms that scour huge data sources to find even shreds of evidence that we can put together to formulate a hypothesis as to what is going on inside a patient. Of course, the doctor has other tools at his/her disposal and is the final judge, but our information can direct them to an answer much faster.
Additional Services
N.A.G.'s algorithms and methodologies are applicable for many other "omics" data including epigenomics, transcriptomics, proteomics, and metabolomics. If you are from a healthcare organization or health clinic with potential for significance volume, please contact us on how you can analyze your patients' "omics" data using our technology. This algorithms can predict disease risk, disease progression, and even options for disease treatment!
We also have an analysis that determines how fast your internal biological clock is progressing - this can give an estimate of how long you may live!
Developer Program
Our platform is open to computer programmers who wish to build apps that add value to peoples' DNA profiles. Note: for security and privacy, each user must specifically approve an app before their data can be accessed. If you would like to join our developer program, please contact us.
Contributing to the Community
N.A.G. is active in the open source community. An example is Pgenesniffer, a utility which finds genomic variants that may be misclassified from pseudogenic sequences.